Design Your Experiment
The following section provides step-by-step instructions for designing a Soluble Protein Workflow experiment.
The design of the experiment can be carried out on the Cloud Software ore usign the provided spreadsheet for the standalone system
- Cloud Connected
- Standalone
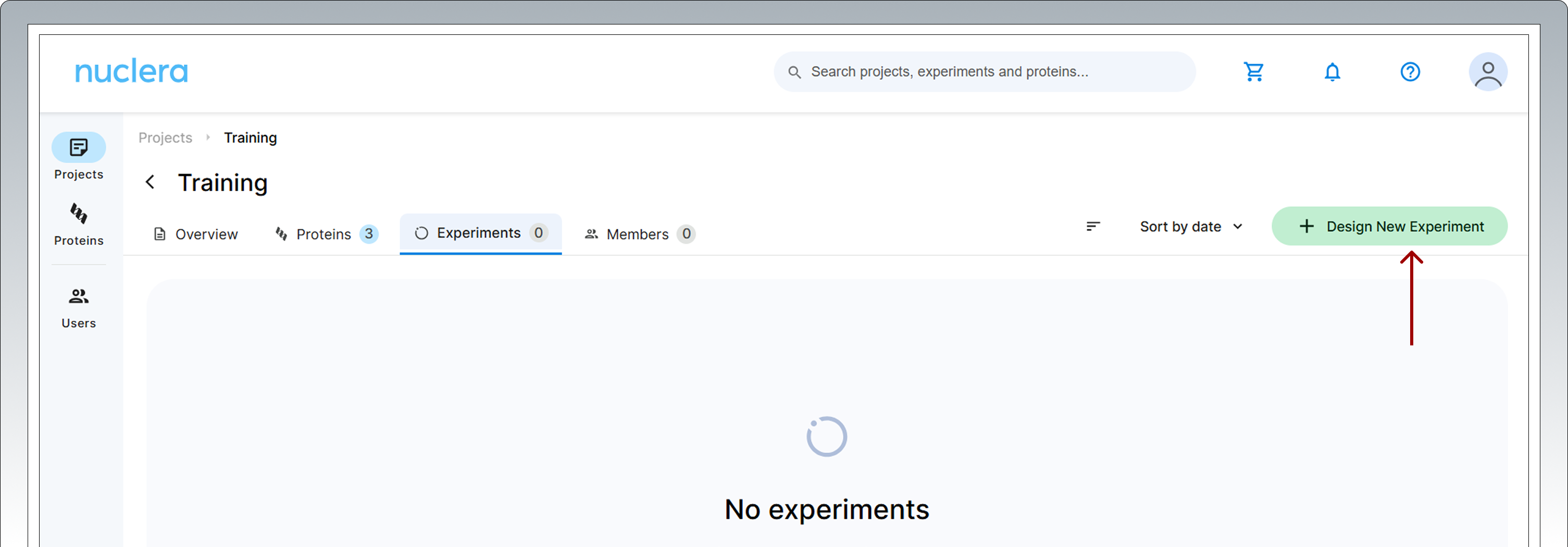
After registering your protein(s), you can move forward to design your experiment. Select the Experiments tab and click on the [Design New Experiment] button.
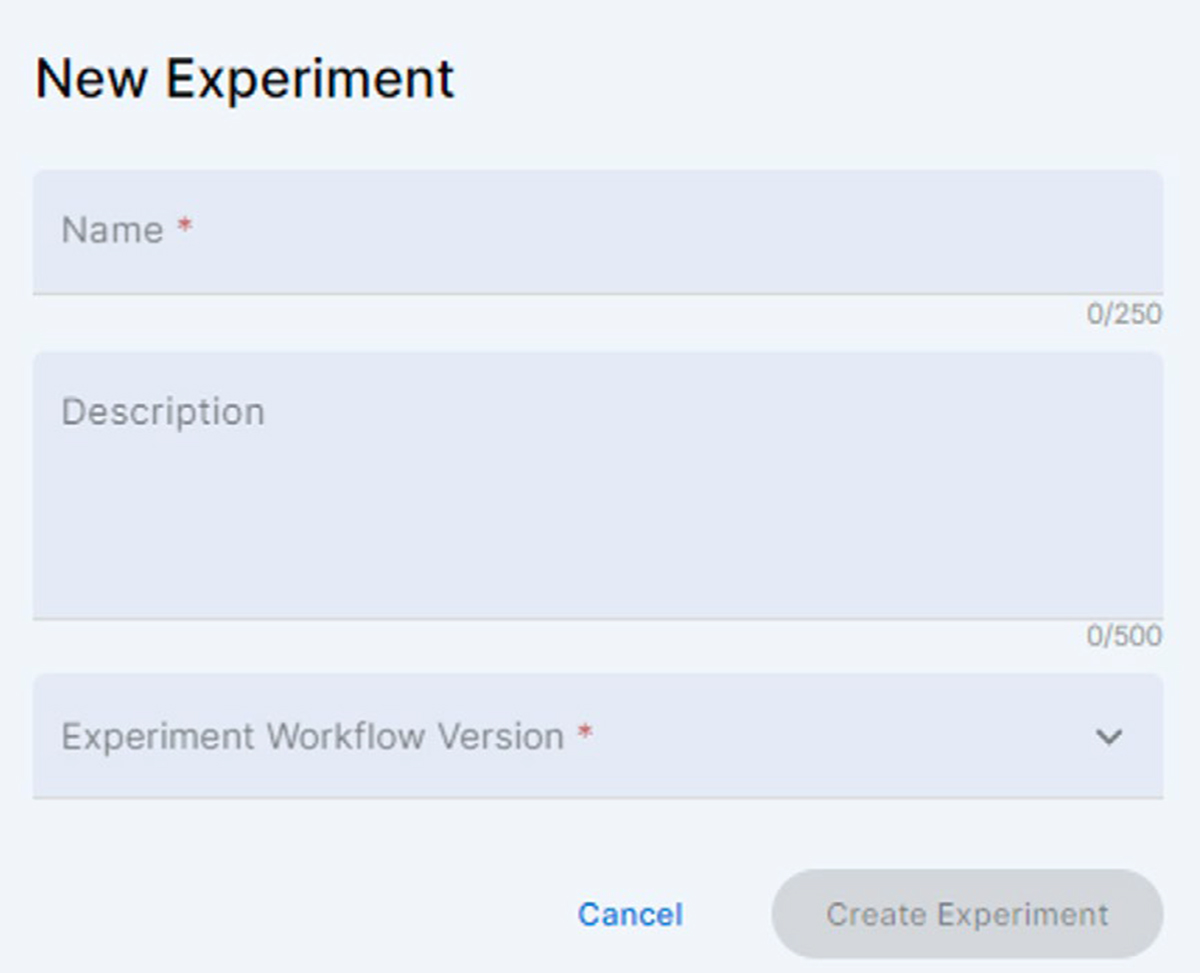
Enter the name of the new experiment, add a short text description and select the Soluble Protein Screen. Click the activated [Create Experiment] button
If the version of the Workflow is not compatible with the current version of the Instrument Software, a warning message is displayed, inviting the user to update the Instrument Software.
Note On the instrument, an incompatible experiment will appear with a warning sign inviting the user to update the Instrument software.
Select the eGene™ Construct
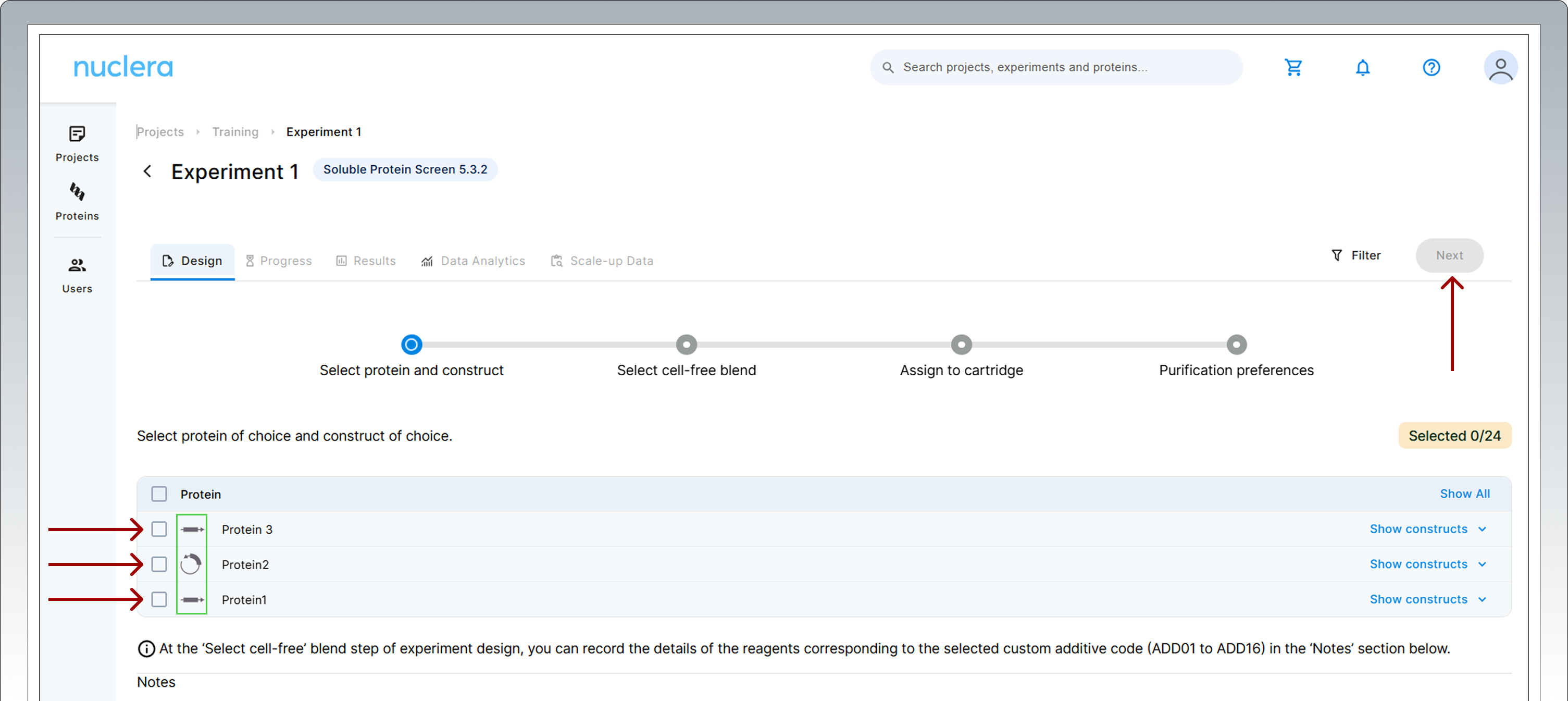
Select 24 DNA constructs to assign to a cartridge. Once a desired number of constructs are selected, click on [Next].
The molecular form of your eGene™ constructs is shown next to the name of the protein chosen (green rectangle).
You can load the same construct (duplicate) in two ports if required by your experimental design.
Select Additives
Select expression conditions by adding two additives to the Cell-free Core Reagent.
If you are in doubt on which additive to choose for your experiment you can refer to the ”Tailored cell free blend” section under “Technology Overview”. If you are undecided on the additive combinations to use, please reach out to Technical Support.
The eProtein Discovery™ system also provides the flexibility to incorporate custom additives into expression and purification workflows, enabling users to tailor conditions for unique protein targets.
To ensure optimal performance and minimize risks, please consult the Compatibility List at https://info.nuclera.com/manual-custom-additives-chemical-compatibility-list.html, which provides detailed guidelines on compatible additives and their maximum allowed concentration. This resource serves as a valuable reference to help you achieve optimal results when working with custom additives. Refer to this before experimenting with custom additives or contact Technical Support if you require more guidance.
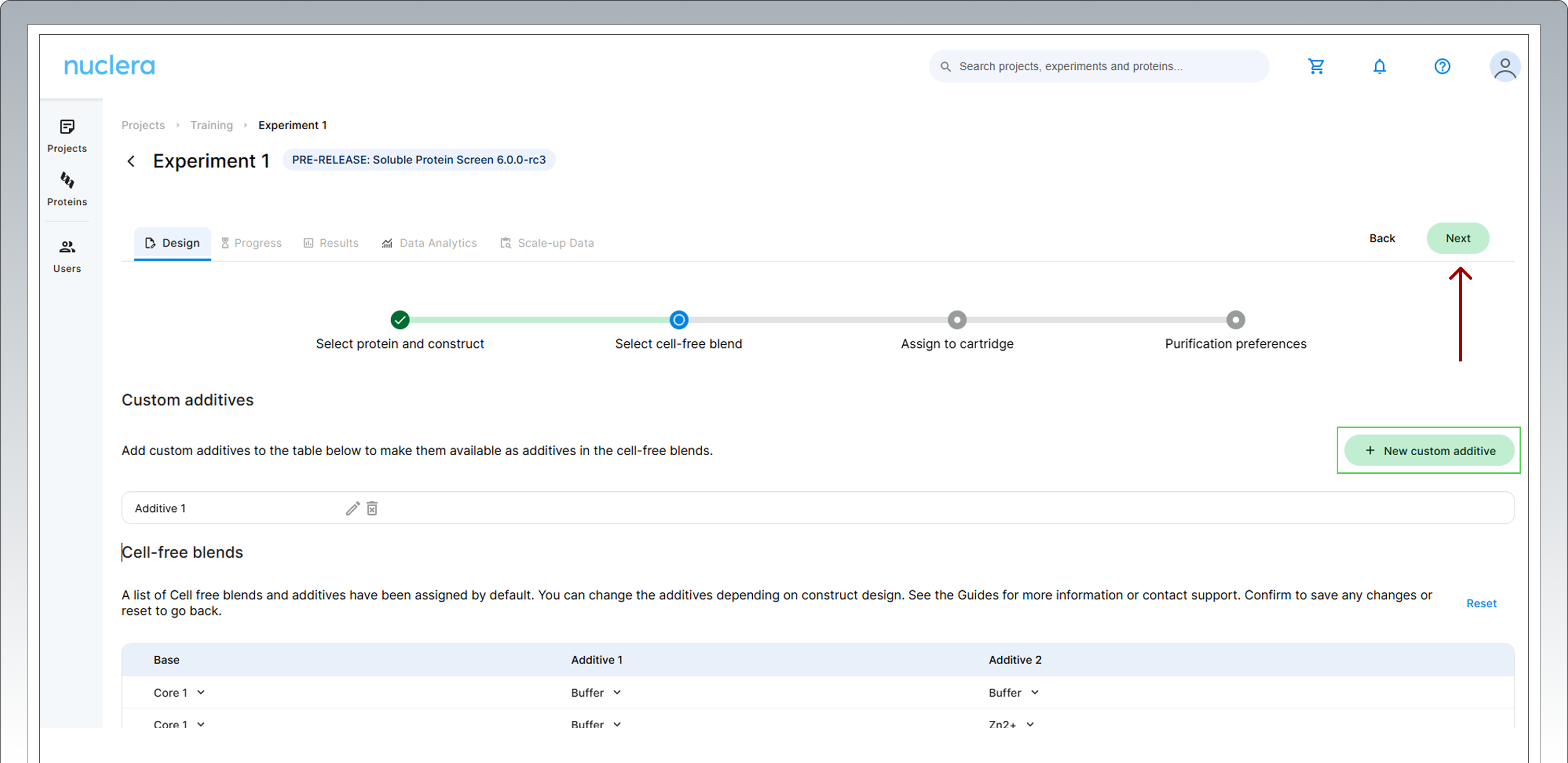
Custom Additives can be shown in the experiment design by clicking on the [New custom additive] box. Up to 20 characters can be used for the definition of the custom additive.
An additive can be selected twice to enhance a specific condition.
Once you finalized the choice of additives to use in the experiment, click [Next] to proceed.
Assign reagents to specific ports
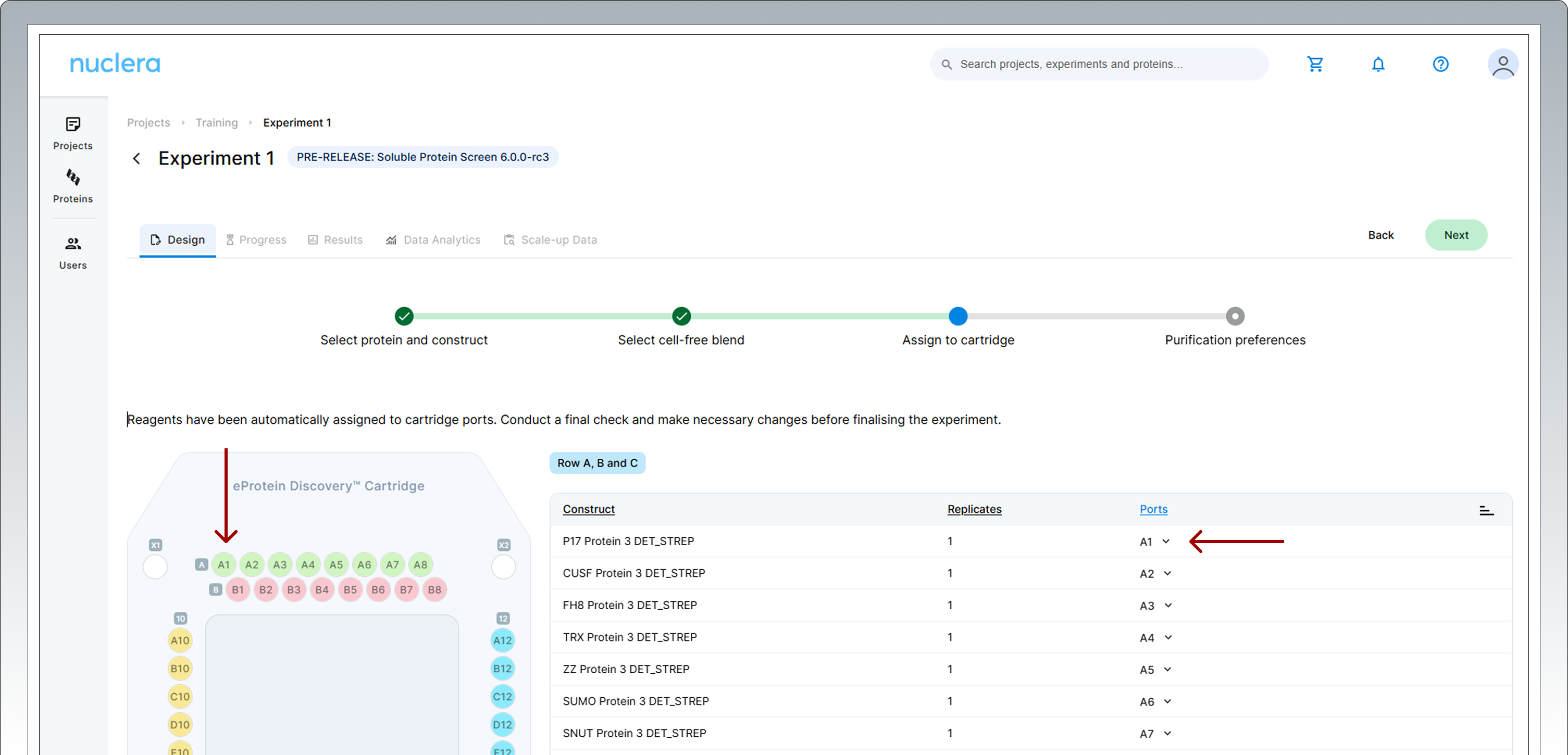
All samples and reagents are now assigned to a specific port on the cartridge.
Review & Correct sample allocation: in the final step you have the opportunity to inspect and potentially change port location for DNA samples and Cell-free Blends.
Drag & drop in the interactive cartridge map or use the port menu available to every sample.
Press the [Next] button to proceed.
Choose Purification Preferences
In the following step you can modify the parameters the instrument will use to select the best expression conditions for purification.
The default variables are pre-selected and are suitable for the majority of experiments.
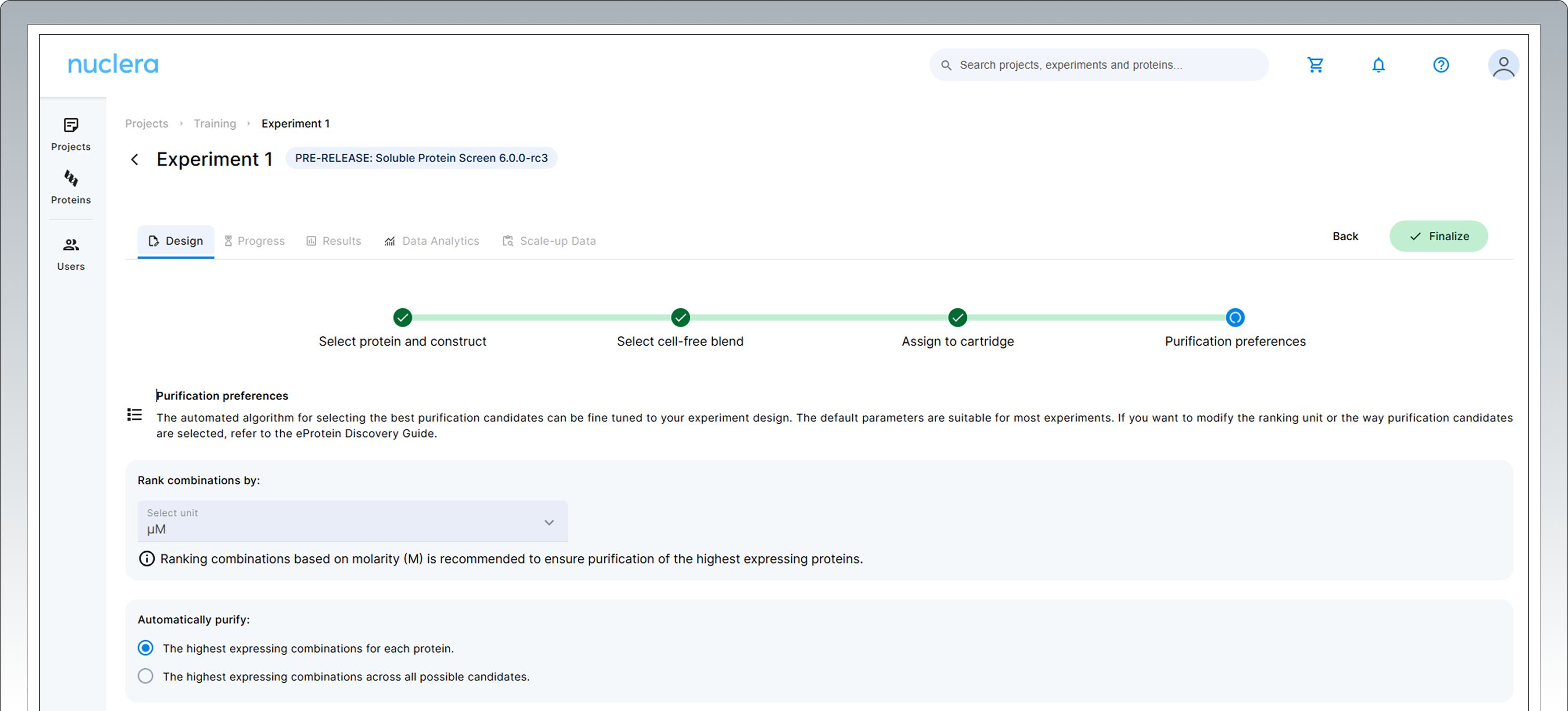
The default algorithm works as follows:
- In total, 30 expression conditions are selected for the purification screen.
- The selection can be done using either molar concentration (µM), recommended, or mass concentration (mg/mL) depending on the purification preferences.
- If you are expressing 3 POI on the cartridge (3 Proteins-of-interest against a 8 fusion tags panel), the instrument will automatically select the 10 highest expressing combinations from each construct so all the constructs are represented in the purification screen (3x10).
- If you are expressing 6 POI (6 POI x4 tags) the instrument will select the 5 highest expressing combinations from each protein (6x5).
As a user you can alter the behavior of the instrument and select "Highest Exprssing combinations across all possible candidates".
Selecting "Highest Exprssing combinations across all possible candidates" can result in some of your protein constructs NOT represented in the 30 conditions selected for purification.
The ranking can be performed using different units, μM or mg/mL. When using mg/mL, higher molecular weight will be favoured. This can result in the construct without any solubility tag to be not selected for purficiation even if they have a better expression (measures as uM )
Finalize the experiment
Press the [Next] button to proceed. Confirm the statements presented by ticking the boxes. If you cannot confirm these four statements we recommend you keep the experiment in draft status (choose [Cancel]).
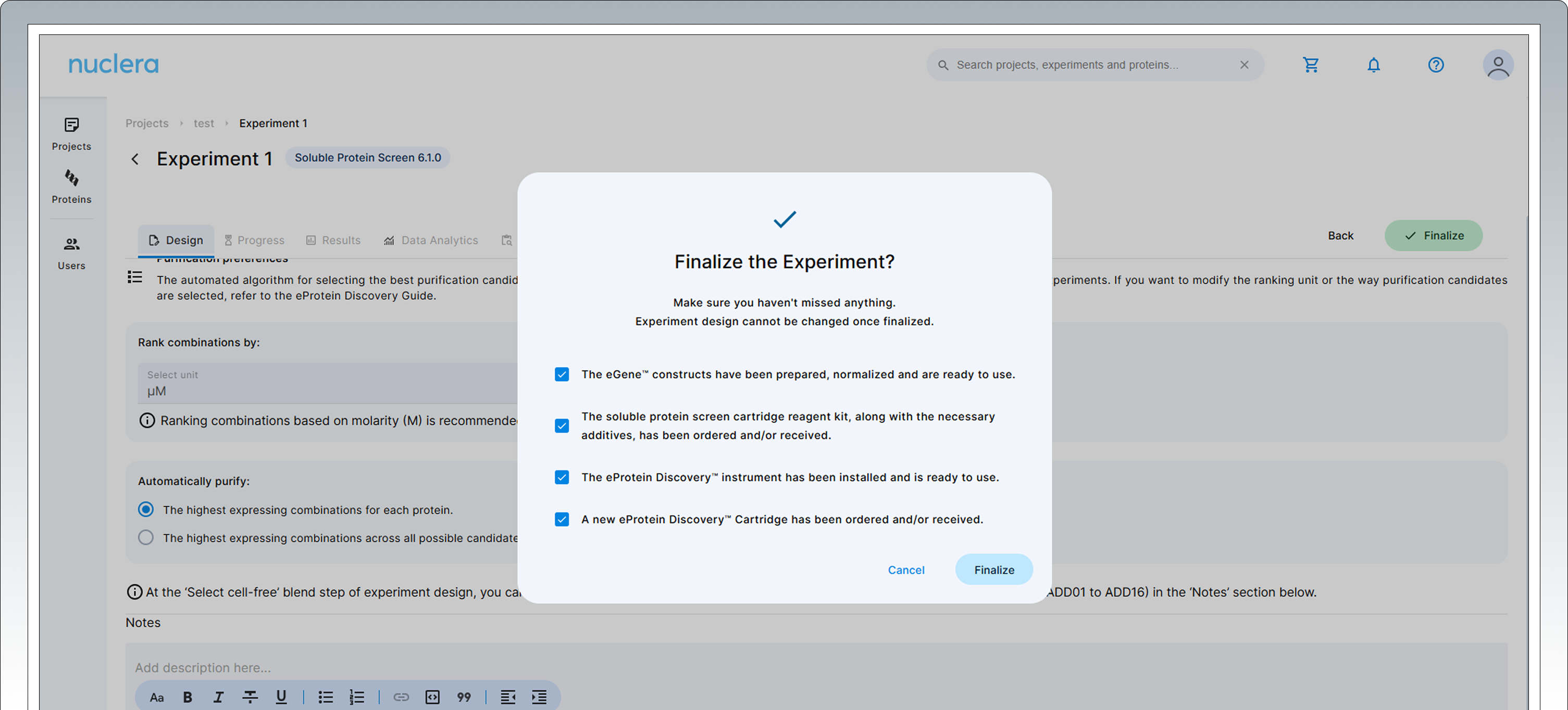
Press the [Finalize] button to proceed. Your experiment is now available on the instrument.
Once finalized, the experimental design cannot be changed.
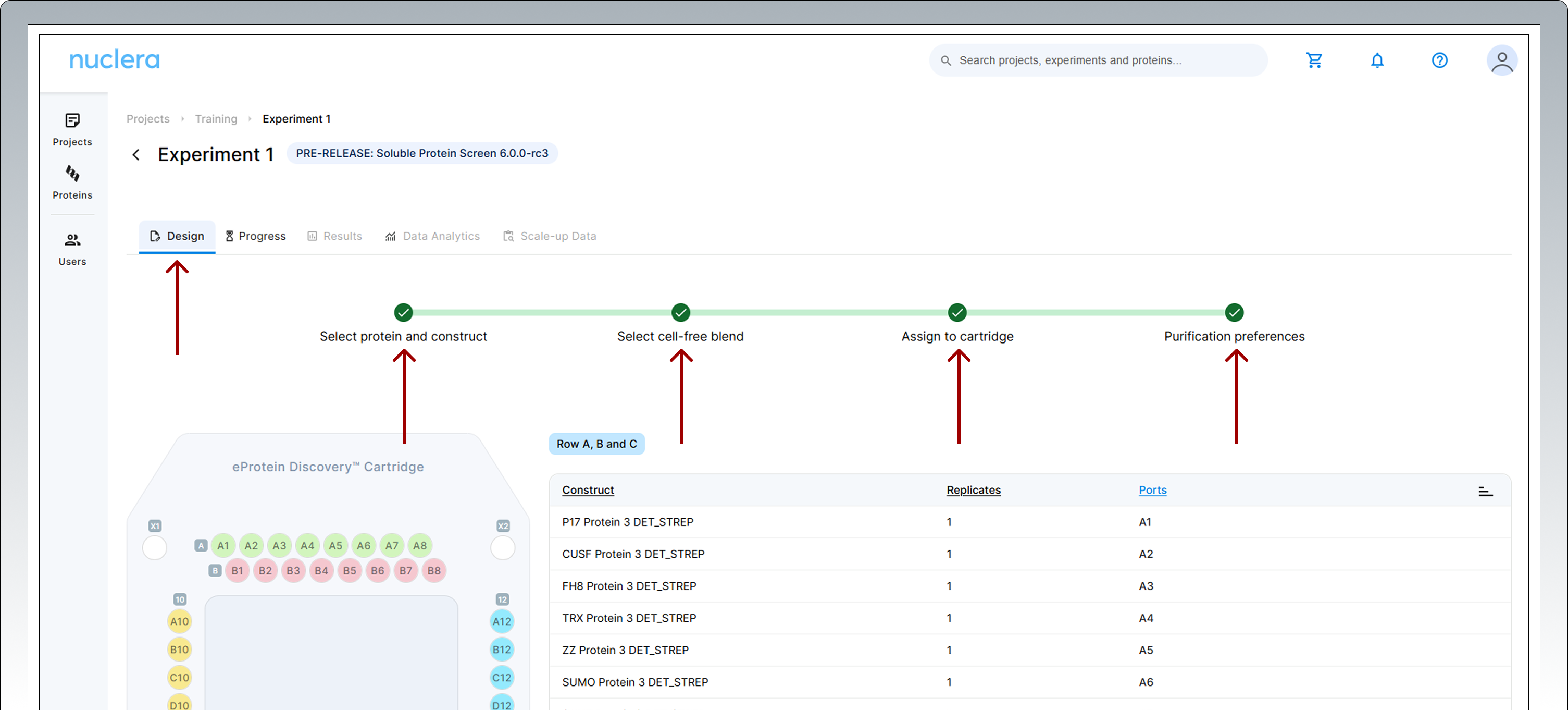
A summary of your experiment is now available. You can toggle and review the various aspects of your experiments by selecting the Design tab and clicking any of the [four green circles] in the design page.
To design an experiment, use the Nuclera eProtein Discovery Standalone template (Excel file) available here, or on request by contacting the Nuclera Technical Support team (techsupport@nuclera.com).
The Nuclera_eProtein_Discovery_Standalone_template file is compatible with Microsoft Excel and it is not compatible with Google Sheet.
The excel template includes multiple worksheets. The worksheets provide step-by-step guide in designing the experiment and analyze data. Read the Template Guidance sheet before proceeding with experiment design.
Open the Excel file, enable editing, and save it under the name of your choice, for example the name or the date of the experiment.
"Template Guidance" sheet
This sheet contains a general introduction and guidance on how to use it. The first steps are to enable editing of the document and to save the file.
"1. Enter Experiment Details" sheet
In this sheet you will design your experiment by adding the combinations of proteins of interest (POI), solubility tags and additives selected for the experiment. Enter the requested information in the cells highlighted in yellow
- Select the format of your experiment (3 proteins x 8 solubility tags, or 4x6 or 6x4 or 24x1)
- Enter the name of the proteins of interest
- Enter the molecular weights (kDa) for each protein of interest
- Select from the drop down menu the solubility tags chosen for the experiment
- Select from the drop down menus the two additives required for your experiment
If you are in doubt on which additive to choose for your experiment you can refer to the “Tailored cell free blend” section under “Technology Overview”. If you are undecided on the additive combinations to use, please reach out to Technical Support.
The eProtein Discovery™ system provides the flexibility to incorporate custom additives into expression and purification workflows, enabling users to tailor conditions for unique protein targets.
To ensure optimal performance and minimize risks, please consult the Chemical Compatibility List at https://info.nuclera.com/manual-custom-additives-chemical-compatibility-list.html, which provides detailed guidelines on compatible additives and their maximum allowed concentration. This resource serves as a valuable reference to help you achieve optimal results when working with custom additives.
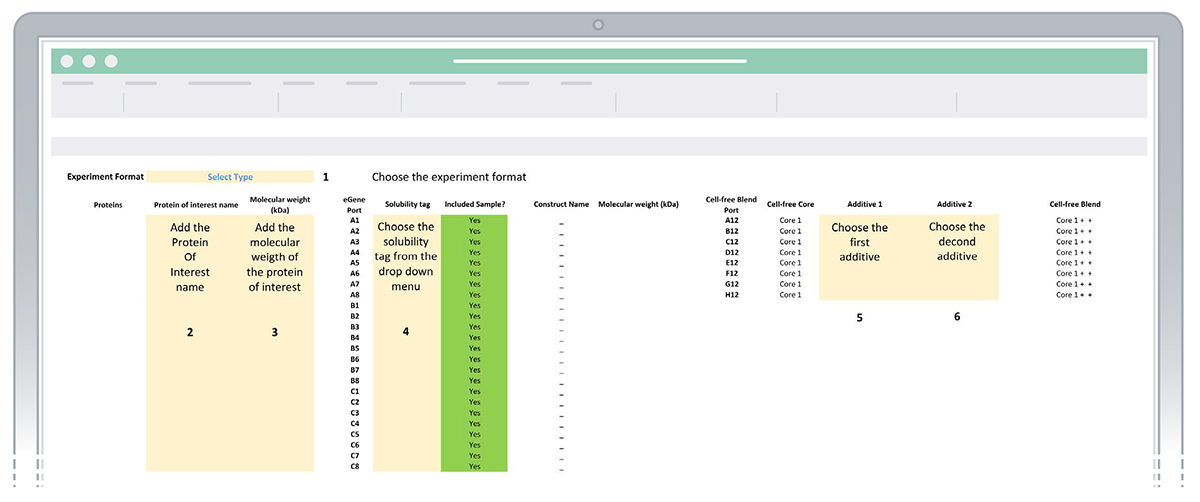
Sheet “1. Enter experiment details” of the eProtein Discovery Standalone template.
- Once the five steps outlined above are completed, the white columns for Construct, Molecular weight (KDa), Protein, and Cell-free Blend will be automatically populated with more information.
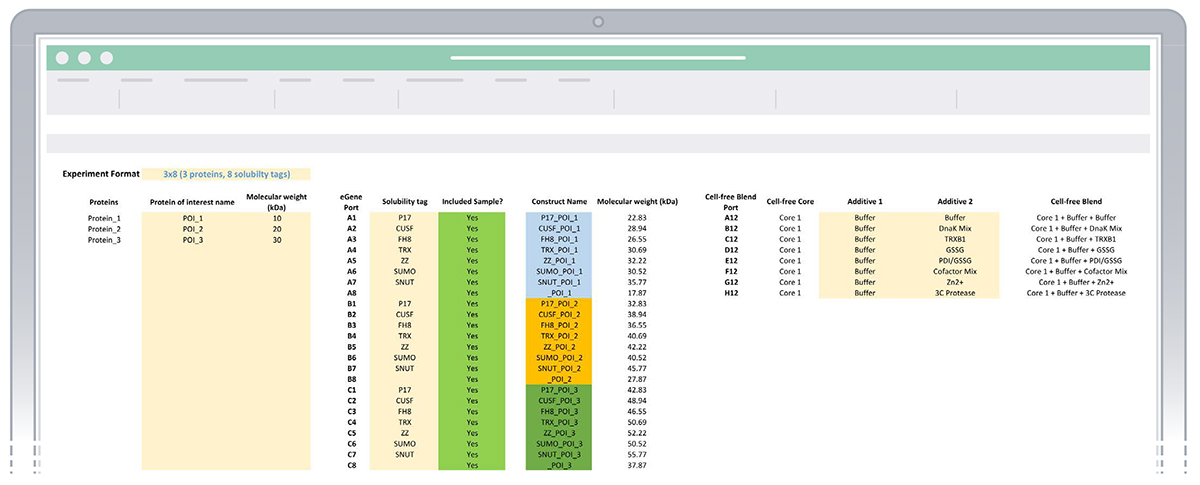
Example of sheet “1. Enter experiment details” with experiment details entered.
"2. Print plate map" sheet
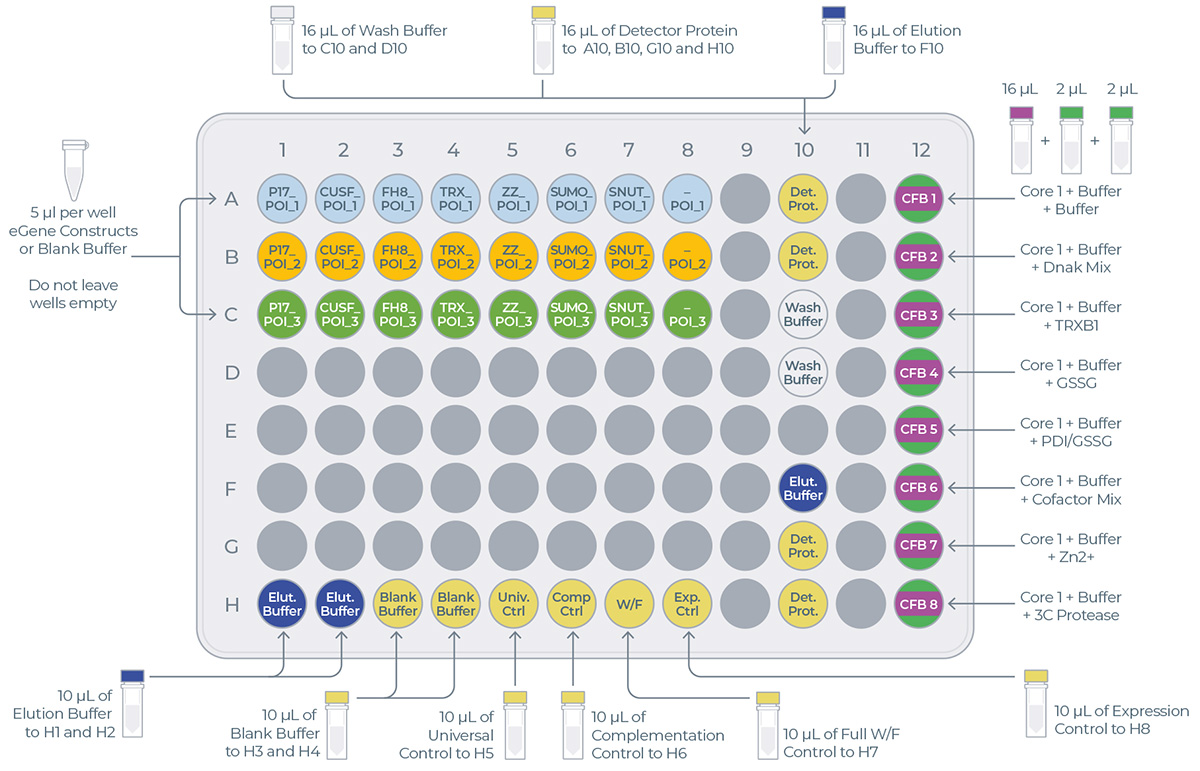
This sheet is the printable version of the experiment design. It is recommended to print this sheet and take it to the lab as a guide for loading reagents onto the transfer plate.
- On the same sheet, below the table, you can find the transfer plate design (Figure 4). This design will help you determine where to load reagents on the transfer plate.
Connect the vial of base fluid to the pump module
In anticipation of starting a new experiment, take a fresh vial of base fluid, open it, and connect it to the left holder on the eProtein Discovery instrument pump module.
It is important to equilibrate the base fluid with the lab atmosphere prior to use. This is to prevent outgassing of the base fluid during the run, as air bubbles can interfere with the droplet movement. We recommend attaching the base fluid to the instrument the day before you will perform the run. An acceptable alternative is to incubate the uncapped base fluid at 30°C / 86°F for 1 hour.
Connect the waste container, empty, to the right holder of the pump module.
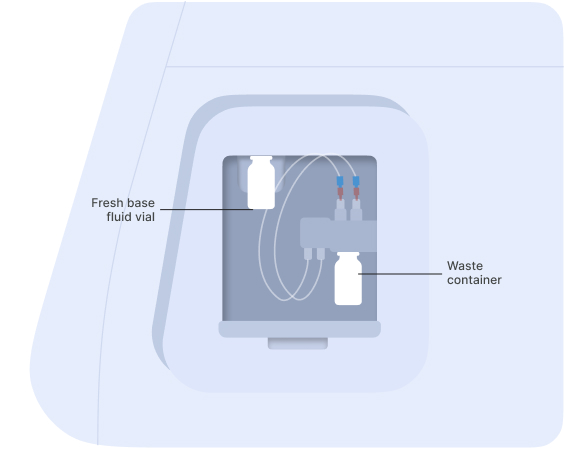
Vial of base fluid and the waste container connected to the pump as shown on the screen
[1]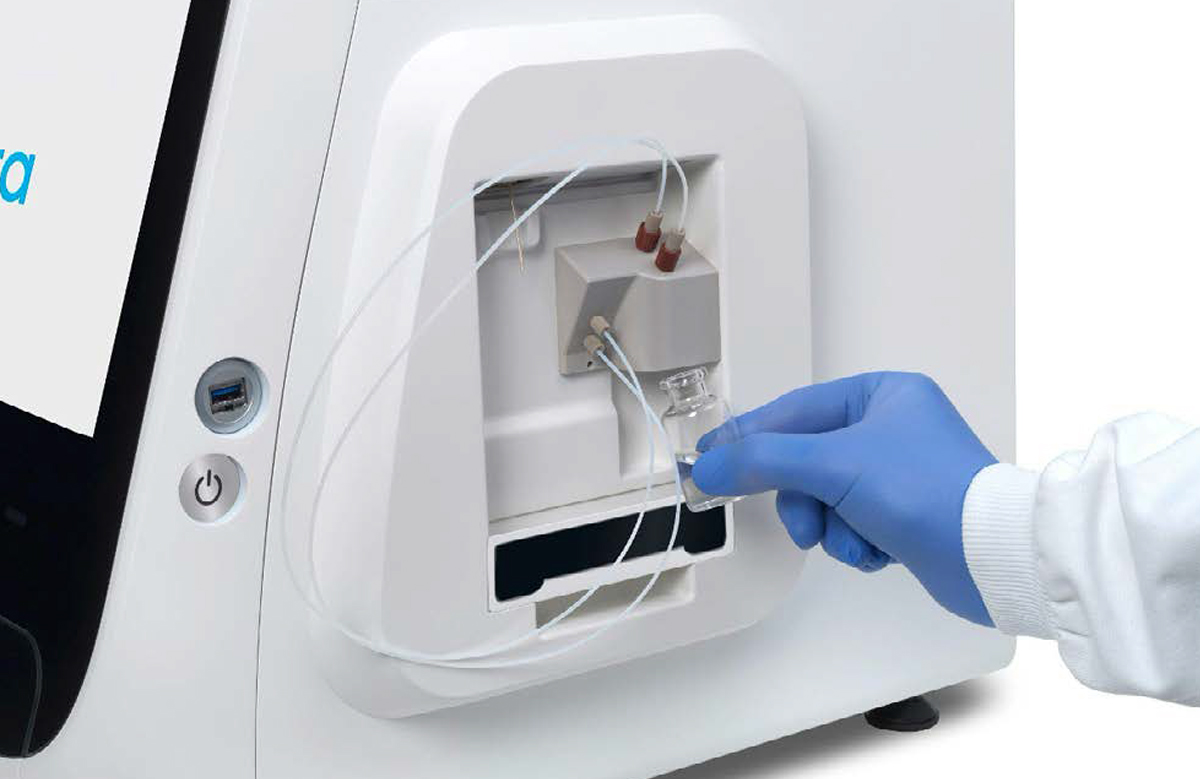 [2]
[2]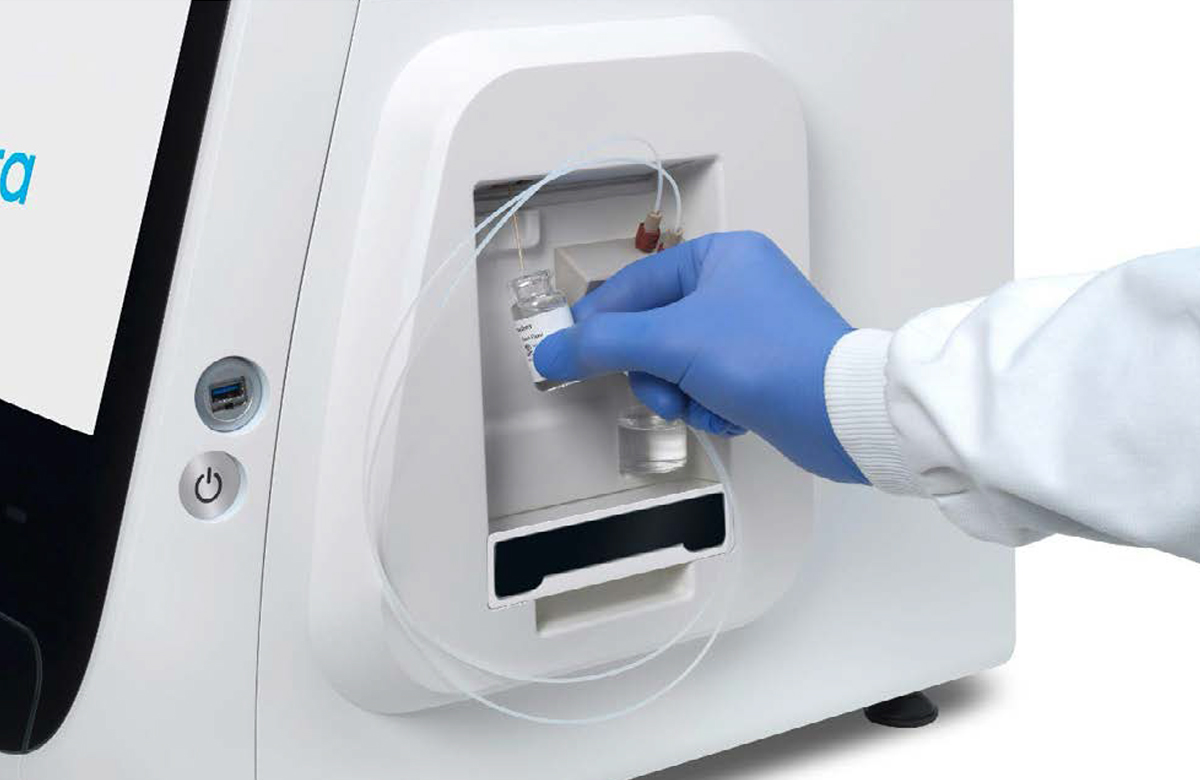
Connection of the empty waste container [1] and the vial of base fluid [2] to the pump